2021
- Khan S, Dunphy A, Anike MS, Belperain S, Patel K, Chiu NH, Jia Z. Recent Advances in Carbon Nanodots: A Promising Nanomaterial for Biomedical Applications. [review] Int. J. Mol. Sci. 22(13), 6786-6817 (2021).
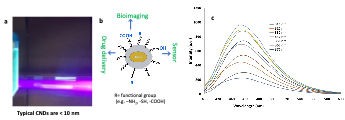
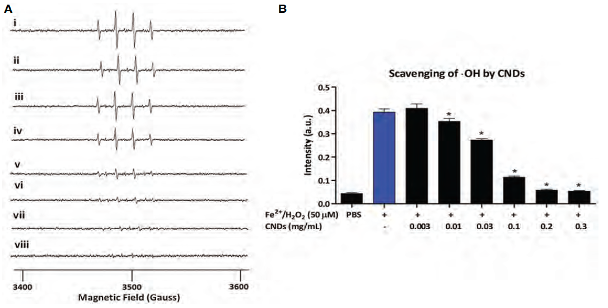
- Khan S, et al. Carbon Nanodots Inhibit Oxidized-LDL-Induced Injury and Monocyte Adhesion to Endothelial Cells through Scavenging Reactive Oxygen Species. J. Biomed. Nanotechnol. 17, 1-14 (2021).
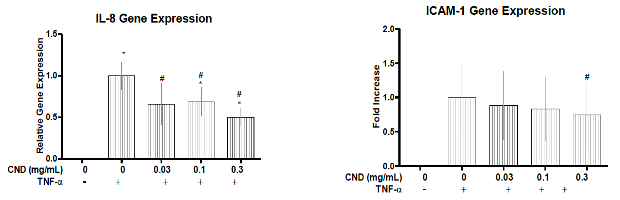
- Belperain S, Kang ZY, Dunphy A, Priebe B,Chiu NH, Jia Z. Anti-inflammatory Effect and Cellular Uptake Mechanism of CarbonNanodots in Human Microvascular Endothelial cells. Nanomaterials 11(5), 1247-1264 (2021).
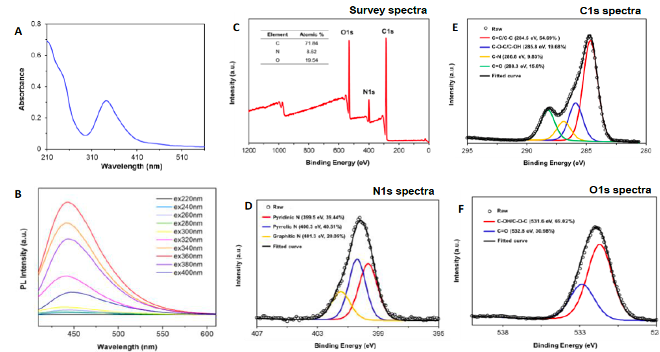
- Dunphy A, Patel K, Belperain S, Pennington A,Chiu NH, Priebe B, Yin Z, Zhu X, Tian S,Wei J, Yi X, Jia Z. Modulation of Macrophage Polarization by Carbon Nanodots and Elucidation of Carbon Nanodot Uptake Routes in Macrophages. Nanomaterials 11(5), 1116-1131 (2021) (Editor’s Choice).
- Chiu NH*, Simpson JH, Wang H, Tannous BA. A theoretical perspective of the physical properties of different RNA modifications with respect to RNA duplexes. BBA Advances, 1, 100025, (2021).

- Wang H, Simpson J, Kotra M, Zhu Y, Wickramsinghe S, Mills D, Chiu NH*. Epitranscriptome of Lactobacillus agilis and its adaptation to growth on inulin. BMC Res. Notes 14, 154-159 (2021).

2020
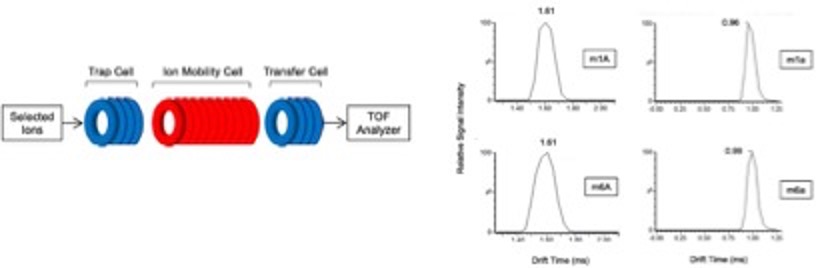
- Wang H, Todd DA, Chiu NH*. Enhanced differentiation of isomeric RNA modifications by reducing the size of ions in ion mobility mass spectrometric measurements. J. Anal. Sci. Technol. 11, 46-56 (2020).
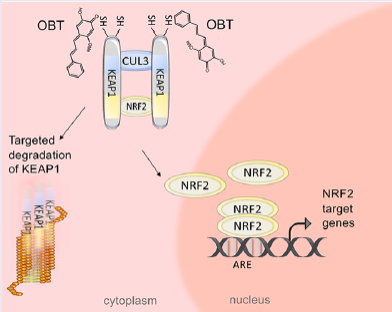
- Badr C, et al. Obtusaquinone is a cysteine modifying compound that targets Keap1 for degradation. ACS Chem. Biol. 15(6), 1445-1454 (2020).
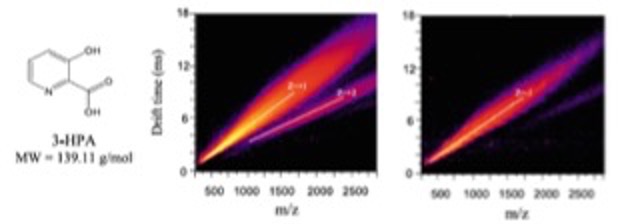
- Mwangi J, Todd D, Chiu NH*. MALDI matrix cluster ions as internal references for ion mobility measurements. Int. J. Ion. Mobil. Spec. 23, 61-67 (2020).
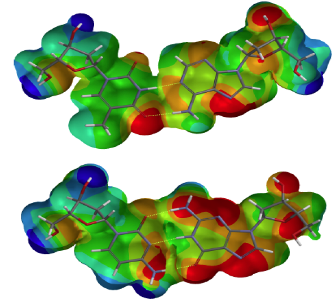
- Proctor NK, Ertan-Bolelli T, Bolelli K, Taylor EW, Chiu NH, Bowen JP. Towards a Better Understanding of Computational Models for Predicting DNA Methylation Effects at the Molecular Level. [review] Curr. Top. Med. Chem. 20(10), 901-909 (2020).
2018
- Mwangi J, Todd D, Chiu NH*. Evaluating the variation of ion energy under different parameter settings in traveling wave ion mobilitymass spectrometry. Int. J. Ion. Mobil. Spec.21(3), 81-86, (2018).

- Uwakweh A, Mwangi J, Todd D, Jia Z, Chiu NH*. Nanospray desorption electrospray ionization mass spectrometry of untreated and treated probiotic Lactobacillus reuteri cells. Anal. Bioanal. Chem. 410 (18), 4237-45, (2018).
2017
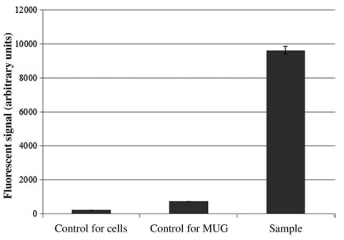
- Chiu NH. Watson A. Measuring β-galactosidase activity in gram-positive bacteria using a whole-cell assay with MUG as a fluorescent reporter. Curr. Protoc. Toxicol. 74, 4.44.1-4.44.8, (2017).
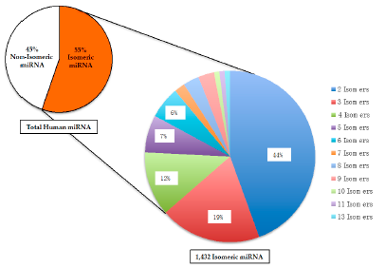
- Cui Z, Chiu NH, Wambua DM. MicroRNA MultiTool: A Software for Identifying Modified and Unmodified Human microRNA Using Mass Spectrometry, Non-coding RNA3(1), 13, (2017).

2016
- Mwangi J, Chiu NH*. High percentage of isomeric human microRNA and their analytical challenges. Non-coding RNA2(4), 13, (2016)
- Watson A, Chiu NH*. Fluorometric cell-based assay for β-galactosidase activity in probiotic gram-positive bacterial cells – Lactobacillus Helveticus. J. Microbiol. Methods 128, 58-60, (2016).

2015
- Watson A, Hu J, Chiu NH*. Single nucleotide polymorphisms in type 2 diabetes among Hispanic population. Diabetes Res. Clin. Pract. 108, e25-27, (2015).

- Wright P, Hu Q, Choi M, Chiu NH, Jia Z. Carbon nanodots interference with lactate dehydrogenase assay in human monocyte THP-1 cells. SpringerPlus 3, 615 (2015).
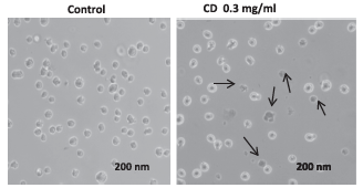
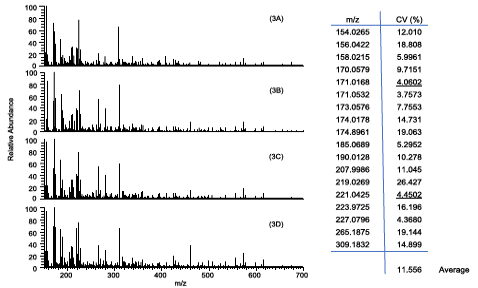
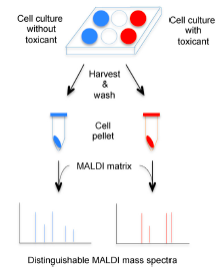
- Chiu NH*, Jia Z, Wright P, Diaz R. Rapid differentiation of in vitro cellular responses to toxic chemicals by using MALDI-TOF mass spectrometry. Environ. Toxicol. Chem. 34(1), 161-166 (2015).
2014
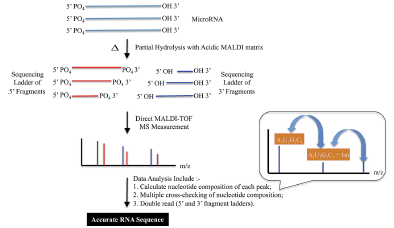
- Wambua D, Ubukata M, Dane J, Cody RB, Chiu NH*. Bottom-up mass spectrometric sequencing of microRNA. Anal. Methods. 6, 8829-8839, (2014).
2012
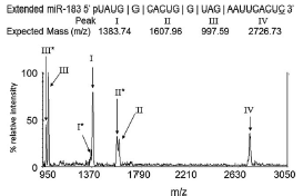
- Wambua D, Tannous BA, Chiu NH*. Creating unique mass signatures for identifying microRNAs. Anal. Methods 4 (10), 3453 – 3459, (2012).
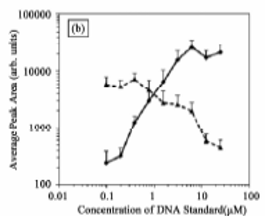
- Chiu NH*, Wilson WB. Unexpected decrease of internal standard signals in quantitative MALDI-TOF MS. J. Anal. Sci. Method Instrum. 2(3), 120-125, (2012).
- Wambua D, Brownstone A, Barnes C, Chiu NH*. Isolation and Detection of Carcinogenic Nucleic Acid Adducts. In “Carcinogen”. Margarita Pesheva, Martin Dimitrov and Teodora Stefkova Stoycheva; InTech: Croatia, 2012; 112-129.
- Chiu NH*, Christopoulos TK. “Advances in Immunoassay Technology”. InTech: Croatia, 2012.
[Downloaded >46,000] - Walleshauser J, Kessler T, Tannous BA, Chiu NH*. A Simple Approach for Evaluating Total MicroRNA Extraction from Mouse Brain Tissues. J. Anal. Sci. Method Instrum., 2(1), 5-12, (2012).
Publications Prior To Tenure
- Zheng X, Xie G, Zhao A, Yao C, Jia W, Chiu NH, Zhou Z, Jia W. The footprints of gut microbial – mammalian co-metabolism. J. Proteome Res. 10(12), 5512-5522, (2011).
- Yang W, Chiu NH. Correlation between UV spectrophotometry and quantitative MALDI-TOF MS. Spectrosc. Lett., 43(7), 602-608, (2010).
- Barnes CA and Chiu NH*. Accurate characterization of carcinogenic DNA adducts using MALDI tandem time-of-flight. Int. J. Mass Spectrom. 279, 170-75 (2009).
- Qing L, Chiu NH and Vouros P. Investigation of benzonase/alkaline phosphatase digestion of DNA and DNA-adducts using LC-ESI MS. Anal. Chem., 79(5), 1907-17 (2007).
- Litman MR, Chiu NH and Wang J. Specific recognition of non-denatured nitrite-oxidizing system of Nitrospira moscoviensis by monoclonal antibody Hyb 153-3. J Chem Technol Biotechnol 81 (3), 318-21 (2006).
- Schiavo S, Yang W, Chiu NH and Krull IS. Comparison of fluorometric detection methods for quantitative PCR. J Immunoassay Immunochem, 26(1), 1-12 (2005).
- Schiavo S, El-Shafey A, Jordan N, Orazine C, Wang Y, Zhou X, Chiu NH, and Krull IS. Immuno-PCR: an overview of the analytical technique currently pushing the limits of detection [review]. PharmaGenomics. January, 36-45 (2004).
- Li G, Zhou X, Wang Y, El-Shafey A, Chiu NH, and Krull IS. Capillary isoelectric focusing and affinity capillary electrophoresis approaches for the determination of binding constants for antibodies to the prion protein. J. Chromatogr A, 1053(1/2), 253-262 (2004).
- Wintzingerode FV, Bocker S, Schlotelburg C, Chiu NH, Storm N, Jurinke C, Van den Boom D. Base-specific fragmentation of amplified 16S rRNA genes and mass spectrometry analysis: A novel tool for rapid bacterial identification. Proc. Nat. Acad. Sci ., 99, 7039 (2002).
- Tannous B, Chiu NH, Christopoulos TK. Heterobifunctional linker between antibodies and reporter genes for immunoassay development. Analytica Chimica Acta, 459, 169 (2002).
- Shahgholi M, Garcia BA, Chiu NH, Heaney PJ, Tang K. Sugar additives for MALDI matrices improve signal allowing the smallest nucleotide change (A:T) in a DNA sequence to be resolved. Nucleic Acids Res., 29; e91 (2001).
- Leushner J, Chiu NH. Automated mass spectrometry: a revolutionary technology for clinical diagnostics [review]. Mol. Diagn. 5; 341(2000)
- White SR, Chiu NH, Christopoulos TK. Expression immunoassay. Methods, 22; 24 (2000).
- Chiu NH*, Tang K, Yip P, Braun A, Koster H, Cantor CR. Mass spectrometry of single-stranded restriction fragments captured by an undigested complementary sequence. Nucleic Acids Res., 28; e31 (2000).
- Chiu NH*, Cantor CR. Mass spectrometry of nucleic acids. Clin. Chem., 45; 1578 (1999).
- Chiu NH, Christopoulos TK. Two-site expression immunoassay using firefly luciferase-coding DNA label. Clin. Chem., 45; 1954 (1999).
- White SR, Chiu NH, Christopoulos TK. Expression immunoassay based on antibodies labeled with a DNA fragment encoding the a-peptide of b-galactosidase. Analyst, 123; 1309 (1998).
- Chiu NH, Christopoulos TK, Peltier J. Sandwich-type DNA hybridization assays based on enzyme amplified time-resolved fluorometry. Analyst, 123; 1315 (1998).
- Chiu NH, Christopoulos TK. Hybridization assays using as a label an expressible DNA fragment encoding firefly luciferase. Anal. Chem., 68; 2304 (1996).
- Christopoulos TK, Chiu NH. Expression Immunoassay. Antigen quantitation using antibodies labeled with enzyme-coding DNA fragment. Anal. Chem., 67; 4290 (1995).

